HomeoSplitter
Disentangling homeologous contigs in allo-tetraploid assembly with homeoSplitter
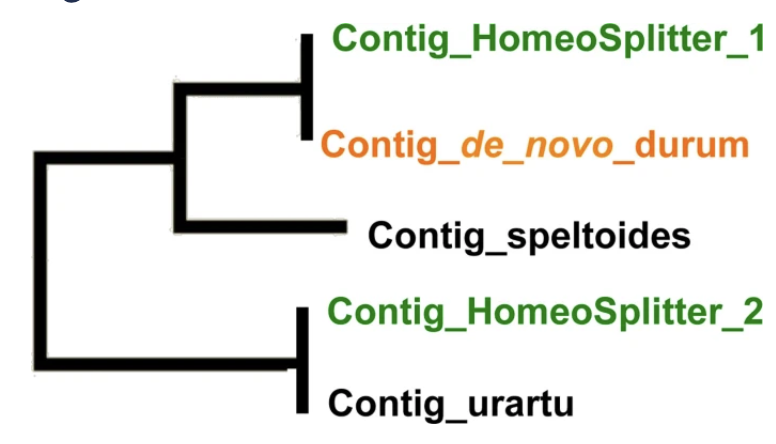
Using Next Generation Sequencing, SNP discovery is relatively easy on diploid species and still hampered in polyploid species by the confusion due to homeology. We develop HomeoSplitter; a fast and effective solution to split original contigs obtained by RNAseq into two homeologous sequences. It uses the differential expression of the two homeologous genes in the RNA.
Running homeoSplitter
HomeoSplitter is a command line program written in Java, it hence runs indifferently on Linux, Mac OS and Windows. To obtain the description of homeoSplitter options, simply execute it without any option:
java -jar ./homeoSplitter_v1.01.jar
You can tune homeoSplitter using its options but only two of them are mandatory (other have default values). The first specifies the input mapping file (in ALR format) while the second specifies the name of the output contigs (in fasta format):
java -jar ./homeoSplitter_v1.01.jar -alrF fic.alr -fastaF seq_after_HS.fasta
The ALR file format
The ALR file, is a text file containing one line per site and one column per accession. For each site the number of occurence of A, C, G and T are denoted between square brackets for each accession.
The easiest way to obtain an ALR file is to map produce a BAM file per accession by mapping its reads on your reference transcriptome. Once these BAM files are sorted and indexed (e.g. using samtools ), you have to create a new text file listing those bam files (e.g. ls *.sorted.bam > BAM_list.txt). You can then use the bam2alr program for converting you BAM files into a single ALR file:./bam2alr -b BAM_list.txt -r input.fasta > output.alr
Handling deletions
Note that both bam2alr and homeoSplitter have options to handle deletion within reads, as a results ALR files will provide the counts for 5 states (A, C, G, T and Del) instead of 4; and contigs output by homeoSplitter may contain deletions (since homeologous copies may differs due to mutation but also due to insertion/deletion events).